Prager R, Rabsch W, Streckel W, Voigt W, Tietze E, Tschäpe H (2003): Molecular properties of Salmonella enterica serotype paratyphi B distinguish between its systemic and its enteric pathovars
J. Clin. Microbiol. 41 (9): 4270-4278.
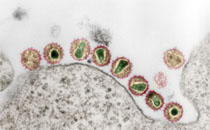
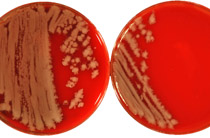
Salmonella enterica serotype O1,4,5,12:Hb:1,2, designated according to the current Kauffmann-White scheme as S. enterica serotype Paratyphi B, is a very diverse serotype with respect to its clinical and microbiological properties. PCR and blot techniques, which identify the presence, polymorphism, and expression of various effector protein genes, help to distinguish between strains with systemic and enteric outcomes of disease. All serotype Paratyphi B strains from systemic infections have been found to be somewhat genetically related with respect to the pattern of their virulence genes sopB, sopD, sopE1, avrA, and sptP as well as other molecular properties (multilocus enzyme electrophoresis type, pulsed-field gel electrophoresis [PFGE] type, ribotype, and IS200 type). They have been classified as members of the systemic pathovar (SPV). All these SPV strains possess a new sopE1-carrying bacteriophage (designated PhiSopE309) with high SopE1 protein expression but lack the commonly occurring avrA determinant. They exhibit normal SopB protein expression but lack SopD protein production. In contrast, strains from enteric infections classified as belonging to the enteric pathovar possess various combinations of the respective virulence genes, PFGE pattern, and ribotypes. We propose that the PCR technique for testing for the presence of the virulence genes sopE1 and avrA be used as a diagnostic tool for identifying both pathovars of S. enterica serotype Paratyphi B. This will be of great public health importance, since strains of serotype Paratyphi B have recently reemerged worldwide.