Kessler JR, Kremer JR, Shulga SV, Tikhonova NT, Santibanez S, Mankertz A et al. (2010): Revealing new transmission routes of measles virus using sequence analysis of phosphoprotein- and haemagglutinin genes
J. Clin. Microbiol.: Epub Nov 24.
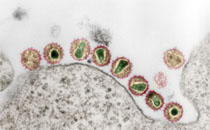
With improved measles virus (MV) control, the genetic variability of MV-nucleoprotein hypervariable region (NP-HVR) decreases. Thus it becomes increasingly difficult to determine the origin of a virus using only this part of the genome. During outbreaks in Europe and Africa we found MV strains with identical NP-HVR sequences. However, these strains showed considerable diversity within a larger sequencing window based on concatenated MV-phosphoprotein and haemagglutinin genes (P/H-pseudo-genes). In Belarus, Germany, Russia and the Democratic Republic of Congo the P/H-pseudo-genes provided insights into chains of transmission, where identical NP-HVR provided none. In Russia, for instance, the P/H-pseudo-gene identified temporal clusters rather than geographical clusters, demonstrating the circulation and importation of independent variants rather than large local outbreaks, lasting for several years, as suggested by NP-HVR. Thus, by extending the sequencing window for molecular epidemiology, a more refined picture of MV circulation was obtained with more clearly defined links between outbreaks and transmission chains. Our results also suggested that in contrast to the P gene, the H gene acquired fixed substitutions that continued to be found in subsequent outbreaks possibly with consequences for its antigenicity. Thus, a longer sequencing window has true benefits both for epidemiological surveillance of measles, as well as for better monitoring of viral evolution.