Lasch P, Drevinek M, Nattermann H, Grunow R, Stämmler M, Dieckmann R, Schwecke T, Naumann D (2010): Characterization of Yersinia Using MALDI-TOF Mass Spectrometry and Chemometrics
Anal. Chem. 82 (20): 8464-8475. Epub Sep 24.
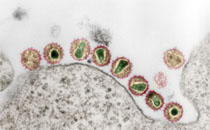
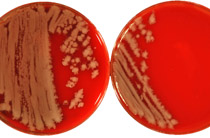
Yersinia are Gram-negative, rod-shaped facultative anaerobes, and some of them, Yersinia enterocolitica, Yersinia pseudotuberculosis, and Yersinia pestis, are pathogenic in humans. Rapid and accurate identification of Yersinia strains is essential for appropriate therapeutic management and timely intervention for infection control. In the past decade matrix-assisted laser desorption ionization time-of-flight (MALDI-TOF) mass spectrometry (MS) in combination with computer-aided pattern recognition has evolved as a rapid, objective, and reliable technique for microbial identification. In this comprehensive study a total of 146 strains of all currently known Yersinia species complemented by 35 strains of other relevant genera of the Enterobacteriaceae family were investigated by MALDI-TOF MS and chemometrics. Bacterial sample preparation included microbial inactivation according to a recently developed mass spectrometry compatible inactivation protocol. The mass spectral profiles were evaluated by supervised feature selection methods to identify family-, genus-, and species-specific biomarker proteins and—for classification purposes—by pattern recognition techniques. Unsupervised hierarchical cluster analysis revealed a high degree of correlation between bacterial taxonomy and subproteome-based MALDI-TOF MS classification. Furthermore, classification analysis by supervised artificial neural networks allowed identification of strains of Y. pestis with an accuracy of 100%. In-depth analysis of proteomic data demonstrated the existence of Yersinia-specific biomarkers at m/z 4350 and 6046. In addition, we could also identify species-specific biomarkers of Y. enterocolitica at m/z 7262, 9238, and 9608. For Y. pseudotuberculosis a combination of biomarkers at m/z 6474, 7274, and 9268 turned out to be specific, while a peak combination at m/z 3065, 6637, and 9659 was characteristic for strains of Y. pestis. Bioinformatic approaches and tandem mass spectrometry were employed to reveal the molecular identity of biomarker ions. In this way, the Y. pestis-specific biomarker at m/z 3065 could be identified as a fragment of the plasmid-encoded plasminogen activator, one of the major virulence factors in plague infections.